GKD: Semi-supervised Graph Knowledge Distillation for Graph-Independent Inference
 Graph Knowledge Distillation
Graph Knowledge DistillationAbstract
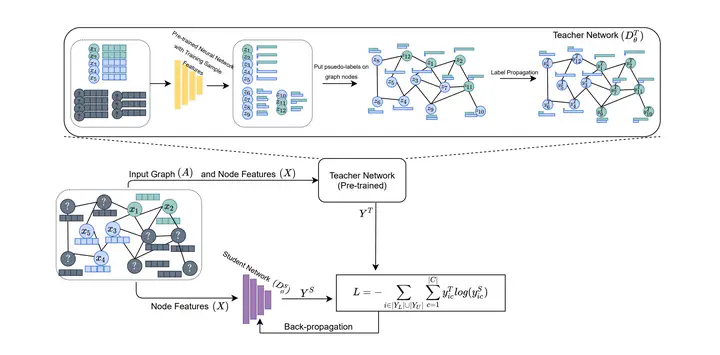
The increased amount of multi-modal medical data has opened the opportunities to simultaneously process various modalities such as imaging and non-imaging data to gain a comprehensive insight into the disease prediction domain. Recent studies using Graph Convolutional Networks (GCNs) provide novel semi-supervised approaches for integrating heterogeneous modalities while investigating the patients’ associations for disease prediction. However, when the meta-data used for graph construction is not available at inference time (e.g., coming from a distinct population), the conventional methods exhibit poor performance. To address this issue, we propose a novel semi-supervised approach named GKD based on the knowledge distillation. We train a teacher component that employs the label-propagation algorithm besides a deep neural network to benefit from the graph and non-graph modalities only in the training phase. The teacher component embeds all the available information into the soft pseudo-labels. The soft pseudo-labels are then used to train a deep student network for disease prediction of unseen test data for which the graph modality is unavailable. We perform our experiments on two public datasets for diagnosing Autism spectrum disorder, and Alzheimer’s disease, along with a thorough analysis on synthetic multi-modal datasets. According to these experiments, GKD outperforms the previous graph-based deep learning methods in terms of accuracy, AUC, and Macro F1.